Introduction
Recent technological developments in fluorescent
microscopy provide new opportunities for multi parameter imaging in living
cells. However, as experiments may run for hours or days, manpower restrictions
apply when controlling and evaluating the experiments. Likewise, the cell’s
sensitivity to phototoxicity creates the necessity to use laser resources
efficiently. This poses a challenge whenever key events happen spontaneously
after hours and then proceed rapidly. Here, overly frequent temporal sampling
(usually chosen at experimental set-up) might lead to premature photoxicity
while too infrequent sampling might result in a poor temporal resolution of the
events under study.
Single cell experiments often follow certain temporal pathways like
biochemical signal transduction with a defined sequence of hallmarks. These
hallmarks are visualized by changes of intensities or distribution of
fluorescent dyes. Different hallmarks may require different set-ups of the
experiment. These set-ups including selected dyes, selected lasers, sampling
speed, spatial magnification are influenced by the biologist’s view on the
underlying process.
Image analysis and automation have been recently used to facilitate
data evaluation in live cell microscopy. High content screening solutions study
large amounts of living cells, use image analysis to detect cell shape
(segmentation) and hence generate data series. Other software packages (Image J,
Cell Tracker) provide cell segmentation and tracking for offline image stacks.
However, we are not aware of any system that allows online evaluation of
temporal intra cellular signals combined with a threshold-based decision system
that adaptively changes the measurement parameter based on a-priori biological
knowledge and is applicable for a wide range of experimental
settings.
Technology
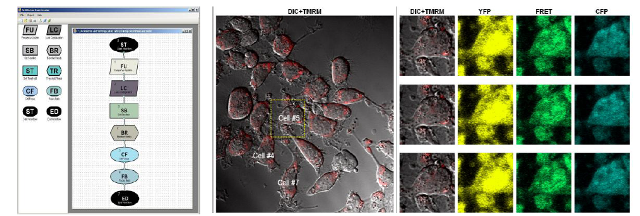
Figure 1:
(Left) The graphical description language for single cell time course
measurement. Top to right presents sequential steps. Lines represent measured
time series data (cells, channels per cell, data series per channel). Boxes
represent image analysis activities, data filtering or parameter setting,
threshold detection or microscope automation. (Center) Cell sample for light
imaging (DIC) and one fluorescent channel (TMRM) for which a threshold is
detected. (Right) Switch to additional fluorescence channels for the labeled
cell at three time points subsequent to threshold detection.
The invention described here uses
image analysis to extract time series of fluorescent signals from cells under
microscopy, compare signal changes within each cell against given thresholds and
subsequently adapt image modalities (sampling rates, laser excitation,
magnification) during single cell measurements by microscope automation. This
thresholding can also be applied to a large range of single cell experiments by
using a graphical language to describe the time course of a single measurement
by defining thresholds and subsequent control actions (see Fig. 1) based on
a-priori biological models of the measurement processes.
Our software system can be integrated into standard fluorescence
microscopes as an analysis and decision logic for performing time lapse single
cell microscopy. Because of the generality of the graphical framework, it can be
applied to a large class of settings ranging from studies in cancer research
(e.g. programmed cell death), cellular physiology, microbiology up to drug
toxicity studies on a high throughput scale.
Applications
The applications of fluorescent imaging are centered on biological
research, such as neuroscience and cancer biology. The technique offers the
ability to image and analyze living cells at higher resolution, which can
provide novel biological insight. Stem cell research is a particularly hot area
of growth.
Advantages
This invention allows an adaptive investigation of single cells
tailored to the studied process (stimulus). It helps to mitigate phototoxicity
since sampling frequency and/or laser resources are increased/switched on
whenever needed, pending on the actual state of the experiment. It reduces
necessary manual intervention like configuration changes in experiments that may
last for several days by microscope automation. As a by-product, this
architecture is aimed to include legacy equipment by decoupling the analysis
process of legacy equipment by separating the microscope specific driver from
the automation and analysis logic
|
Feature
|
Benefit
|
|
Automated
detection of biological events |
Event
resolution at higher sampling rate leading to new biological insight
|
|
Adaptation
of image modalities in response to biological events
|
Reduction of
photo-toxicity and photo-bleaching through economic use of resources
|
|
Online
automated image analysis |
Less effort
than manual analyses |
Contacts:
Dr Aoife Gallagher, RCSI Technology Transfer, 123 St Stephen’s
Green, Dublin 2, Ireland.
Email: aoifegallagher1@rcsi.ie Tel: +353 1
4022394
Dr. Gearóid Tuohy, RCSI Technology Transfer, 123 St Stephen’s
Green, Dublin 2, Ireland.
Email: gearoidtuohy@rcsi.ie Tel: +353 1 4022362
Principle Investigator:
Dr. Heinrich Huber, Royal College of Surgeons, 120 St Stephen’s
Green, Dublin 2, Ireland.
Email:
Heinhuber@rcsi.ie